How do I increase the minimum dot size in Seurat's DotPlot function? · Issue #2297 · satijalab/seurat · GitHub

By A Mystery Man Writer
Thank you very much for your hard work in developing the very effective and user friendly package Seurat. I want to use the DotPlot function to visualise the expression of some genes across clusters. However when the expression of a gene

Negative average expression value on Dotplot · Issue #2379 · satijalab/ seurat · GitHub
SpatialPlot distortion when setting crop=TRUE · Issue #5141 · satijalab/ seurat · GitHub
DotPlot: cluster order and subsets · Issue #711 · satijalab/seurat · GitHub

Average expression of gene on dotplot · Issue #2649 · satijalab/seurat · GitHub

Dot plot scaling issue/request · Issue #396 · scverse/scanpy · GitHub

How to add average expression scale to dotplot of merged gene list (plotted onto single dot plot) · Issue #4544 · satijalab/seurat · GitHub

Dot plot not changing average expression · Issue #3337 · satijalab/seurat · GitHub
Dot plot Metrics · Issue #2921 · satijalab/seurat · GitHub
DotPlot: cluster order and subsets · Issue #711 · satijalab/seurat · GitHub
- Lululemon size dot, what does it mean? Informative

- Different Sized Dots Royalty-Free Images, Stock Photos & Pictures

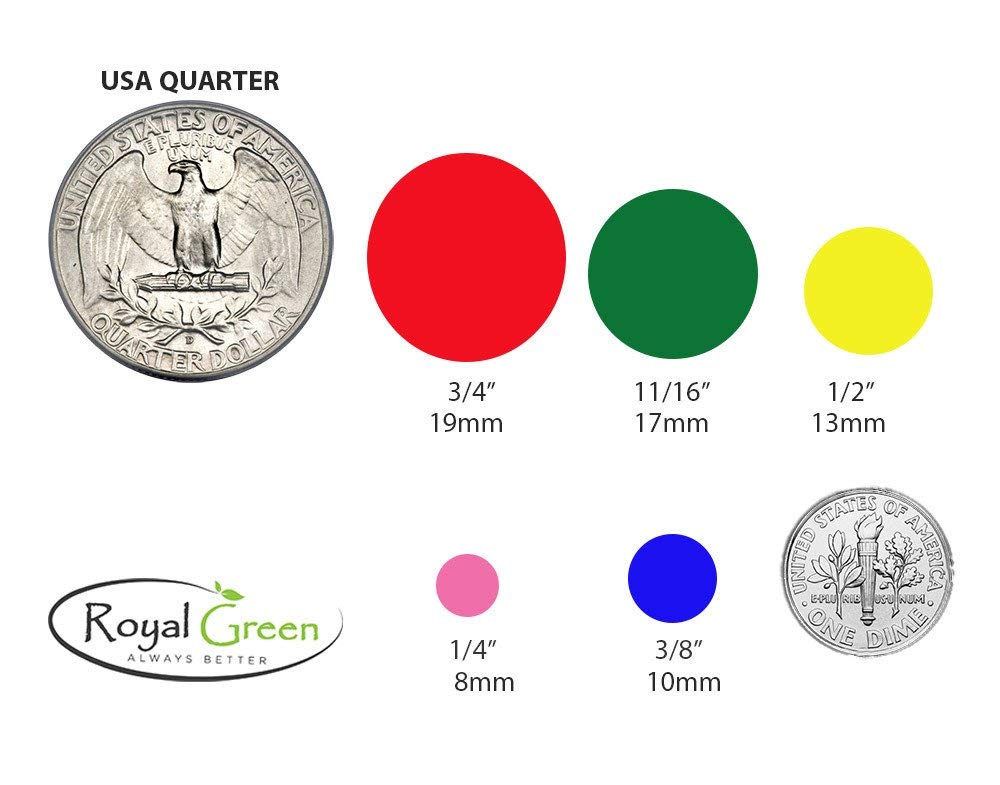
- Colored Dot Stickers for Children's Crafts, Games, and Arts.
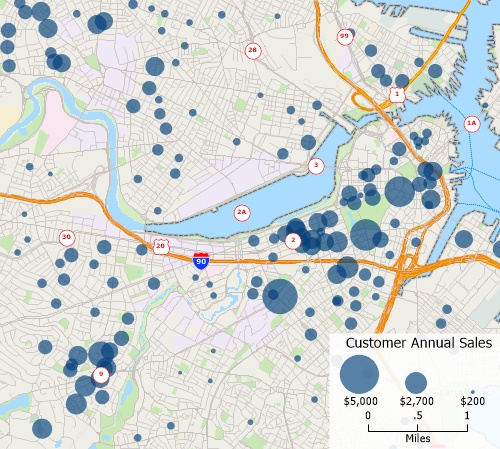
- What is a Thematic Map - Thematic Map Definition
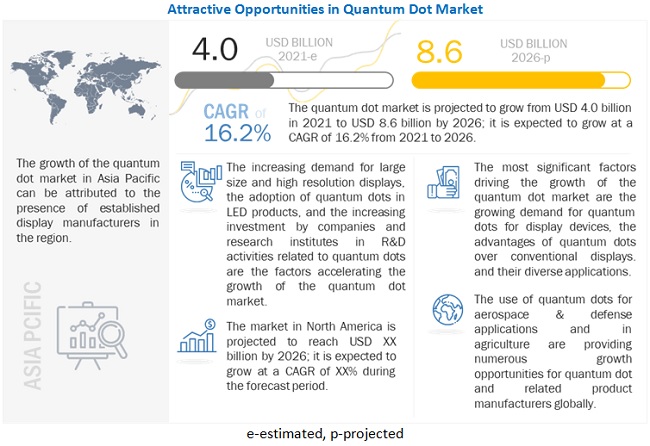
- Quantum Dot Market Size, Share, Industry Trends, Companies, Growth Analysis 2030